Research Articles
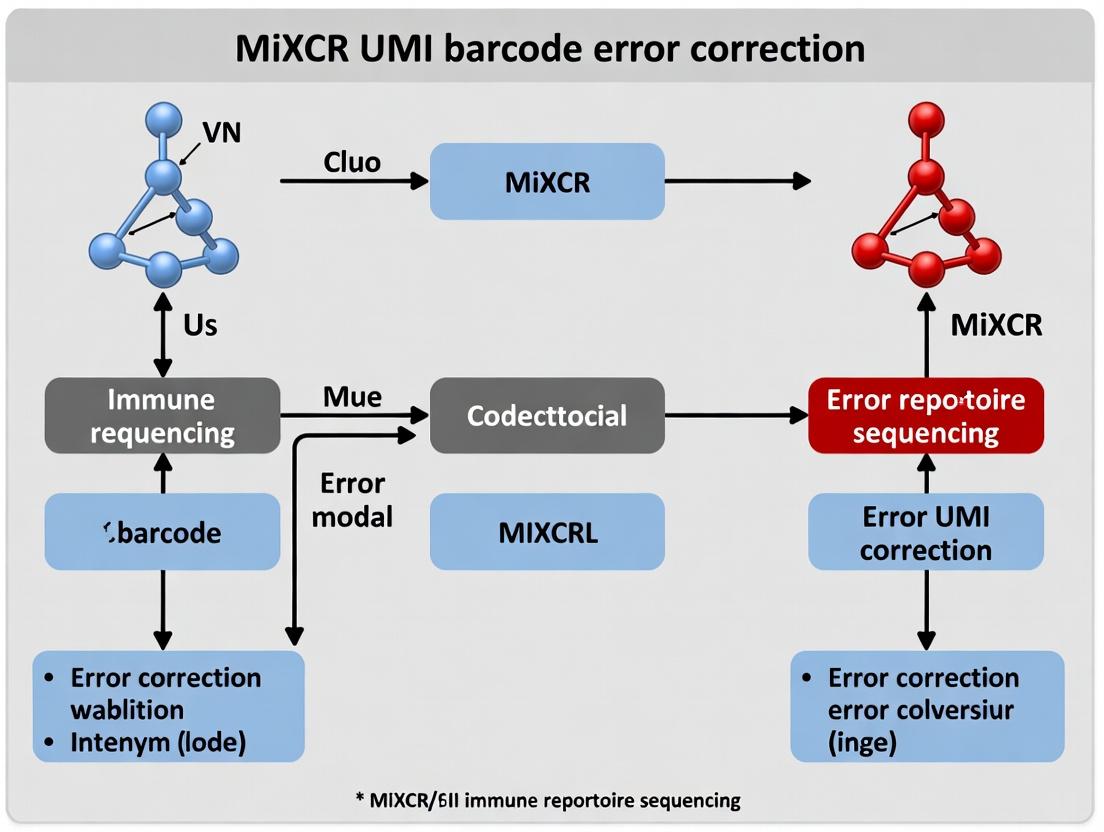
MiXCR UMI Error Correction: A Complete Guide to Accurate Immune Repertoire Profiling
This article provides a comprehensive guide to Unique Molecular Identifier (UMI) error correction within the MiXCR pipeline for immune repertoire analysis.
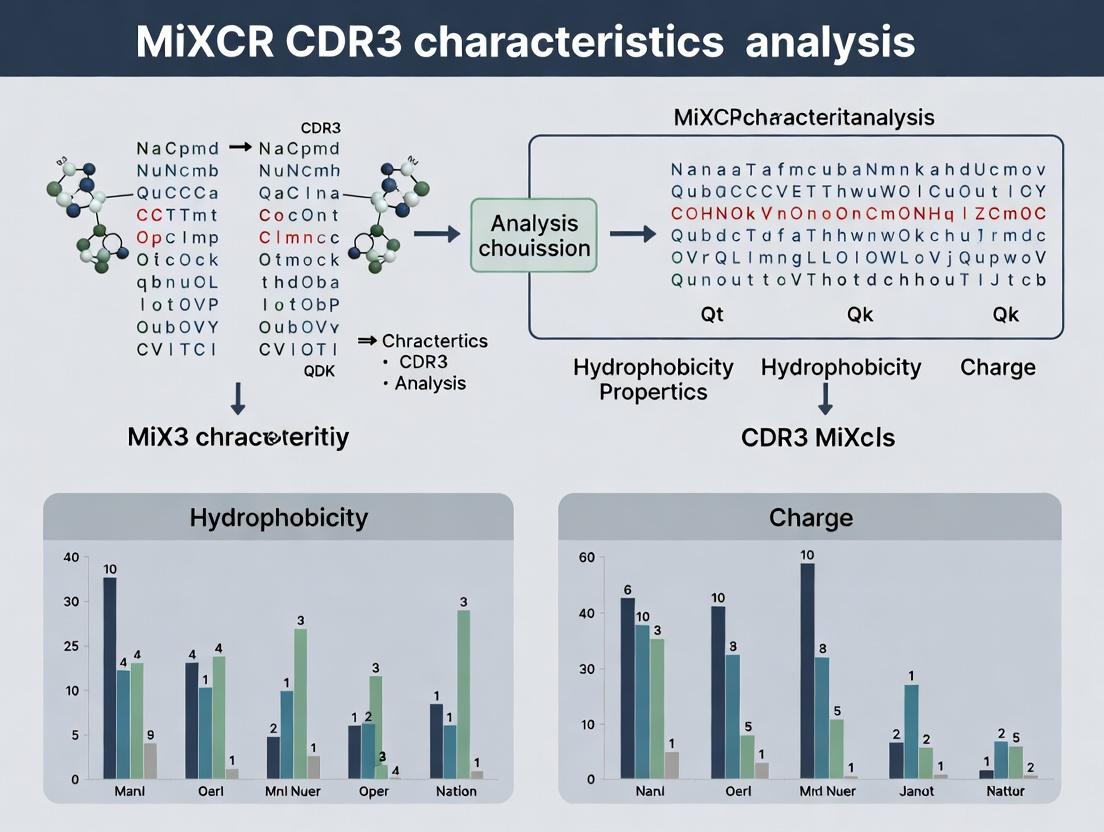
Decoding Adaptive Immunity: A Practical Guide to Analyzing CDR3 Hydrophobicity and Charge with MiXCR
This comprehensive guide empowers immunology researchers and therapeutic developers to extract and interpret critical physicochemical properties of T-cell and B-cell receptor repertoires using MiXCR.
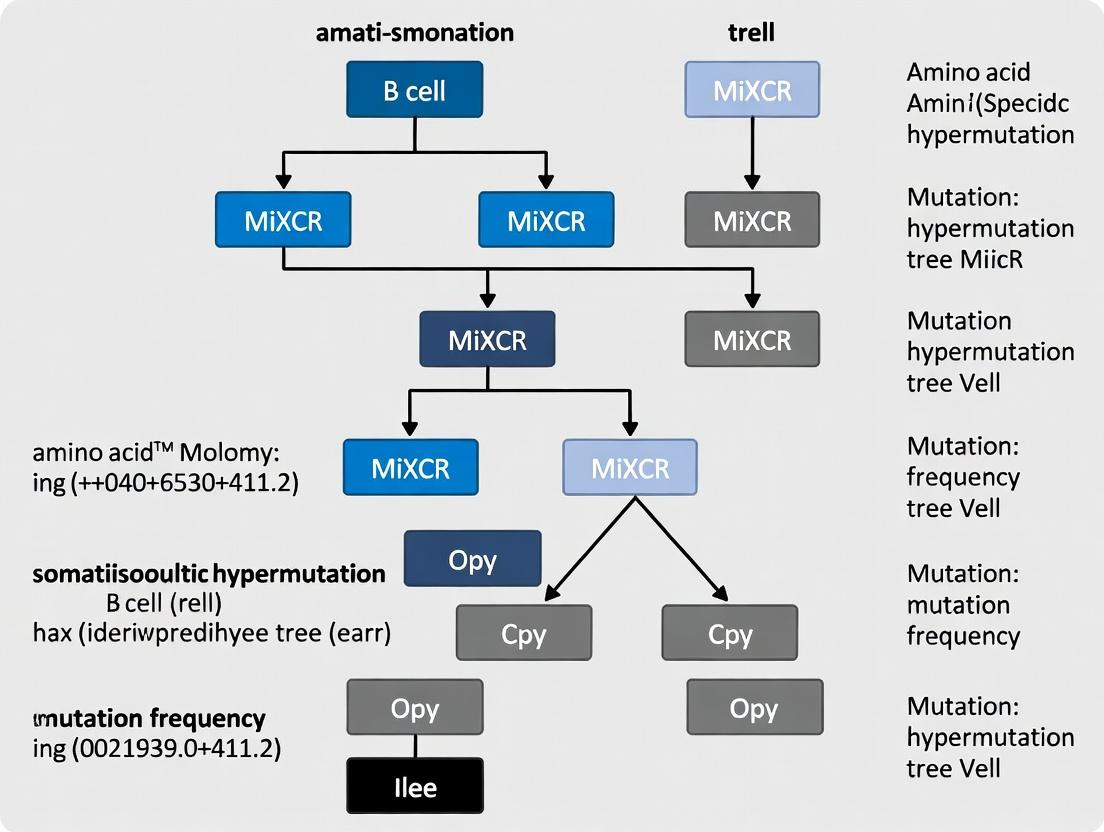
Decoding Antibody Evolution: A Comprehensive Guide to B Cell SHM Tree Analysis with MiXCR
This article provides a detailed guide for researchers, scientists, and drug development professionals on analyzing B cell somatic hypermutation (SHM) lineage trees using MiXCR.
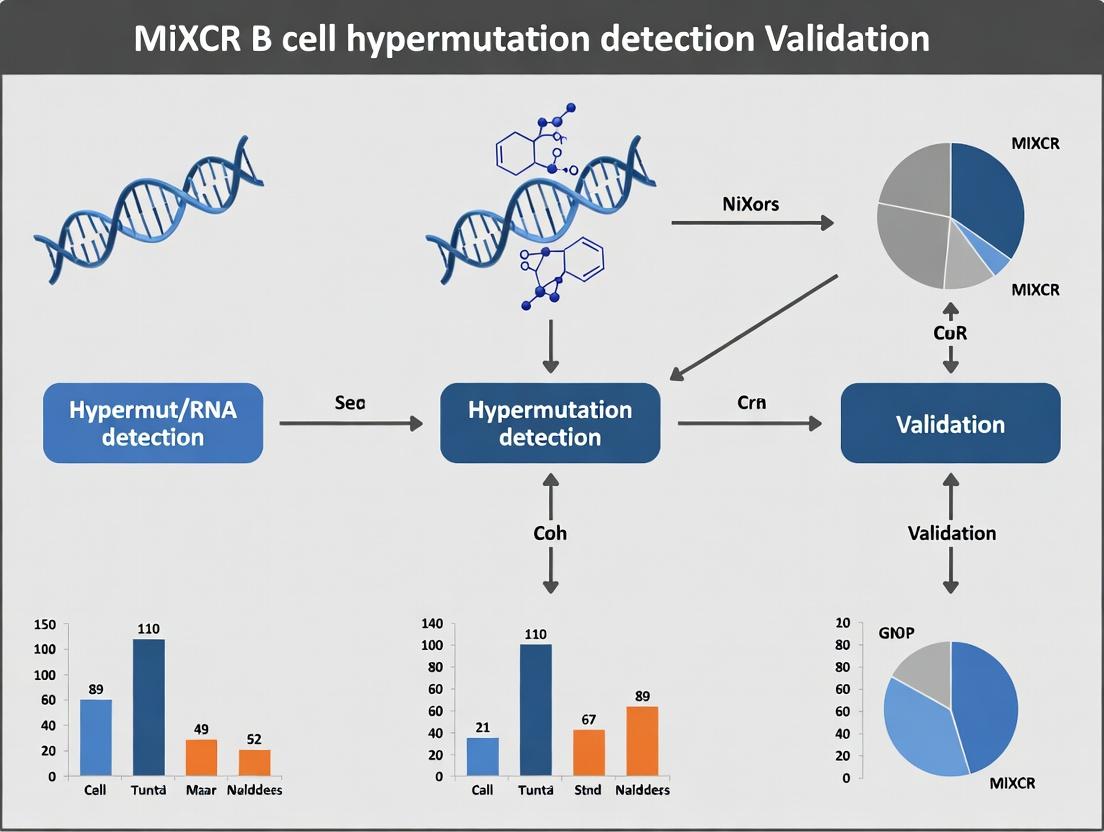
Validating B Cell Hypermutation Analysis with MiXCR: A Comprehensive Guide for Researchers in Immunology and Drug Development
This article provides a detailed guide to validating B cell receptor (BCR) somatic hypermutation (SHM) detection using the MiXCR software suite.
MiXCR AIRR-Compliant Export: A Complete Guide to Format Compatibility for Immunogenomics Research
This comprehensive guide explores MiXCR's compatibility with the Adaptive Immune Receptor Repertoire (AIRR) Community data standards.
Benchmarking Single-Cell Integration Tools: A 2024 Comparative Analysis of MOFA+, Seurat, and Harmony
This article provides a comprehensive, data-driven comparison of three leading single-cell multi-omics integration tools: MOFA+, Seurat, and Harmony.
Scalable Multi-Omics Integration: A Deep Dive into MOFA+ and Stochastic Variational Inference for Large-Scale Biomedical Datasets
This article provides a comprehensive guide to scaling Multi-Omics Factor Analysis (MOFA+) for large datasets using variational inference.
MOFA+: The Ultimate Guide to Multi-Omics Single-Cell Data Integration for Biomedical Research
This comprehensive guide explores MOFA+, a powerful statistical framework for integrating multi-omics single-cell data.
Decoding Sepsis Complexity: A Guide to MOFA+ for Immune Cell Heterogeneity Analysis in Drug Development
This article provides a comprehensive guide for biomedical researchers and drug development professionals on applying Multi-Omics Factor Analysis plus (MOFA+) to dissect immune cell heterogeneity in sepsis.
MOFA+, Seurat, LIGER, and GLUE: Benchmarking Integration Power for Single-Cell Multi-Omics Analysis in 2024
This article provides a comprehensive, up-to-date performance comparison and practical guide for four leading single-cell multi-omics integration tools: MOFA+, Seurat (v5), LIGER, and GLUE.